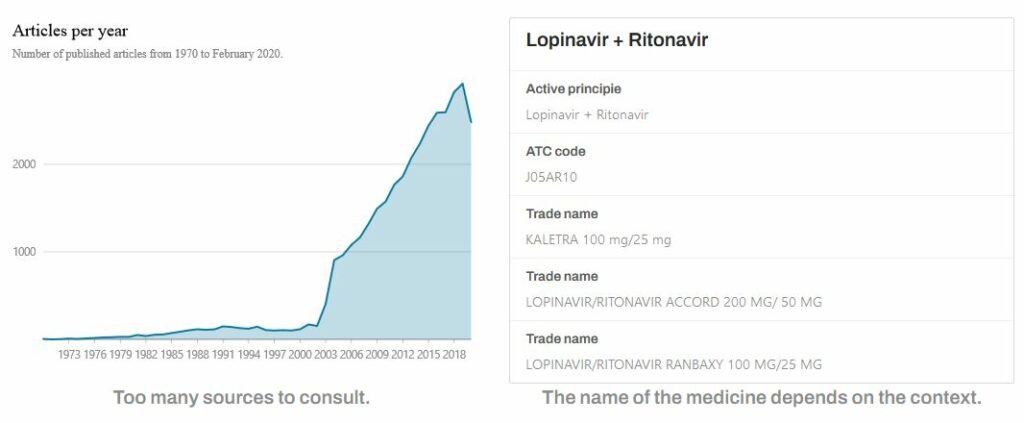
The idea for this project was born during the early months of the pandemic. In March and April 2020, hospital admissions for COVID-19 created a drug supply problem. In response, it became necessary to research the scientific literature on the disease to find available alternative drugs.
The published scientific literature, describing the active ingredients, drugs and their use, proved to be as numerous as difficult to manage, to the point that the White House Office of Science and Technology Policy appealed to the international Artificial Intelligence community, opening its CORD-19 repository of scientific articles related to the coronavirus. It is a repository of more than 300,000 scientific documents in English, with full-text access to more than 200,000 of them.
The Ontology Engineering Group of the Universidad Politécnica de Madrid, with a successful track record in the areas of Ontological Engineering, Semantic Web and Linked Data, Natural Language and Semantic e-Science, contributed by launching the Drugs4Covid project.
The CORD-19 corpus has been the main database used in this study. Each week a new update is published that augments the initial collection with new publications. The April 2020 edition, with 60,702 scientific articles and 2,103,891 paragraphs, was the first used in the study. The latest indexed update is the January 2022 edition, with 334,580 scientific articles and more than 10 million paragraphs.
All this content is being analyzed using natural language processing and knowledge extraction techniques through a 4-step process: (1) Process and index the corpus of scientific papers, and all of its paragraphs by identifying active ingredients, therapeutic groups, symptoms and diseases. (2) Automatically annotating, using Artificial Intelligence techniques, each text with this information, and requesting the help of citizen scientists to (3) review these annotations and propose relationships between them. Finally, (4) publishing a knowledge graph with all the data and their evidences, that is, connecting and unifying the information in a meaningful way, to make it interrogable in a natural way through applications that facilitate the search and navigation. In order to facilitate access to the information collected in the knowledge graph, a question-answer interface is being developed that allows queries to be made in natural language and returns answers also in natural language. orem ipsum dolor sit amet, consectetur adipiscing elit. Ut elit tellus, luctus nec ullamcorper mattis, pulvinar dapibus leo.
All of these results are being published as open data so they can be used by the healthcare community and any other research team interested in solving additional problems. In fact, the language models created to identify drugs, diseases and genes in the scientific literature had more than 100,000 downloads on HuggingFace in its first month of publication.
More information on the project can be found on the project’s website: https://drugs4covid.oeg.fi.upm.es/